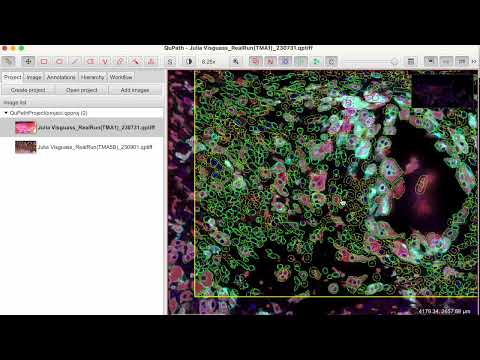
The script collection for QuPath.
You don't need to install the script.
While some scripts can be executed as commands,
most of them must be run within the QuPath GUI.
The cell segmentation requires the cellpose and stardist extension in QuPath.
git clone [email protected]:jianhong/codex_qupath.git
## list available scripts
ls codex_qupath/scr
## list available stardist models
ls codex_qupath/models/stardist
This step involves merging the results from Cellpose and Stardist analyses.
First, you'll need to install the Cellpose and Stardist extensions. Don't forget to set the Python path for the Cellpose extension.
Next, navigate to line 25 of the cellpose_stardist_multiNucleis_underGUI.groovy script and update the path of the stardistModel variable to match the file path of your cell_seg_model.pb. Adjust the parameters from line 26 to line 38 accordingly. You may want to change the cytoChannels and measurementPercentileCutoff according to your observations. The distanceCutoff is the maximal cutoff distance of nucleus for multiple nucleus cells.
Then, select a region in the opened TIFF view window in QuPath. Start with a small region when testing the script, and expand it once you achieve the desired results.
Finally, run the cellpose_stardist_multiNucleis_underGUI.groovy script by clicking the Run button in the Script Editor.
The script ExportCellDetectionMeasurement.groovy is designed to export cell segmentation data. Upon export, two files are generated and saved in a folder prefixed with "measurementsExport".
- The
.tsvfile is compatible withcreateSeuratObj.R. - The
.geojsonfile is intended for reloading intoQuPath.
These exported files contain comprehensive information, including cell area, locations, marker signal statistics, and nucleus classification. The cell locations are particularly useful for neighborhood analysis.
The script fill_detections.groovy is employed to duplicate detected cells into Detections and Annotations objects within QuPath. Subsequently, user can establishe the classes in QuPath to assign colors.
Users may want to try different classifier for the cell type detection by reset the line10-25. Here I set an example for the classifier.
The scripts were originally a collection from published paper, Image.sc Forum, and imagescientist.com or written by Jianhong Ou.
This work were supported by Duke Regeneromics and Visgauss Lab @Duke.edu.
If you would like to contribute to this collection, please create a pull request.
For further information or help, don't hesitate to get in touch on issue channel.
If you use jianhong/codex_qupath for your analysis, please cite it using the following url: https://github.com/jianhong/codex_qupath.
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.