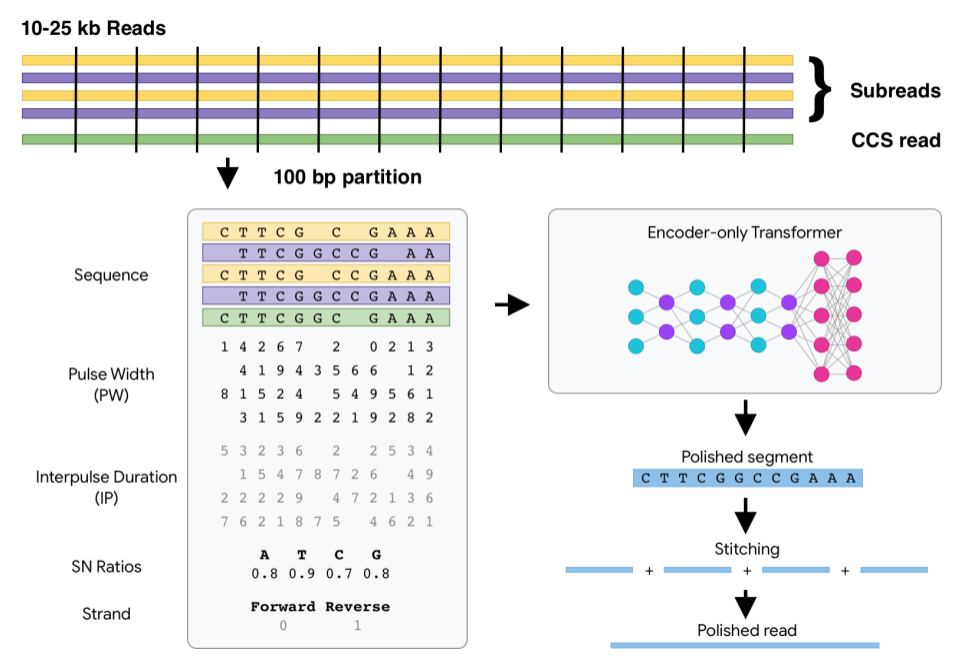
DeepConsensus uses gap-aware sequence transformers to correct errors in Pacific Biosciences (PacBio) Circular Consensus Sequencing (CCS) data.
pip install deepconsensus==0.1.0You can ignore errors regarding google-nucleus installation, such as ERROR: Failed building wheel for google-nucleus.
git clone https://github.com/google/deepconsensus.git
cd deepconsensus
source install.sh(Optional) After source install.sh, if you want to run all unit tests, you can
do:
./run_all_tests.shSee the quick start.
After a PacBio sequencing run, DeepConsensus is meant to be run on the CCS reads and subreads to create new corrected reads in FASTQ format that can take the place of the CCS reads for downstream analyses.
See the quick start for an example of inputs and outputs.
NOTE: This initial release of DeepConsensus (v0.1) is not yet optimized for speed, and only runs on CPUs. We anticipate this version to be too slow for many uses. We are now prioritizing speed improvements, which we anticipate can achieve acceptable runtimes.
If you are using DeepConsensus in your work, please cite:
DeepConsensus: Gap-Aware Sequence Transformers for Sequence Correction
This is not an official Google product.
NOTE: the content of this research code repository (i) is not intended to be a medical device; and (ii) is not intended for clinical use of any kind, including but not limited to diagnosis or prognosis.